Alyssa Culver, Dr. Jesse Barber

Background
Bats and moths have been in an evolutionary arms race for at least 50 million years. To get a better understanding our lab has gone out around the world, recording thousands of different moths ultra sound anti-bat clicks (Barber & Conner 2007).
Methods
- Collected acoustic data from moths around the world
- Created a file name pipeline
- Clipped moth acoustic files for each individual moth
- Three analyzed cycles
- One 100 millisecond cycle
- One 350 millisecond cycle
- Create spectrograms of clipped files
- Analyzed spectrograms using a machine-learning approach to Python.
Data
Work in progress as still clipping files and creating spectrograms
AMMO BAR
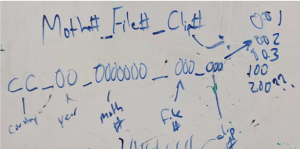
Keep file names relative to the data that was collected, but in such a way that each file name is unique.
Decipher strong clicking patterns amongst weak patterns or bat calls
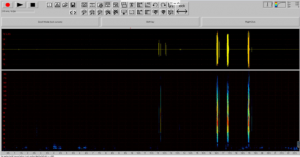
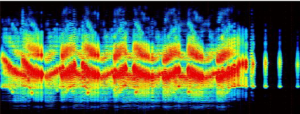
Developing a pipeline for a meta database will make future data collection more readily available for analysis.
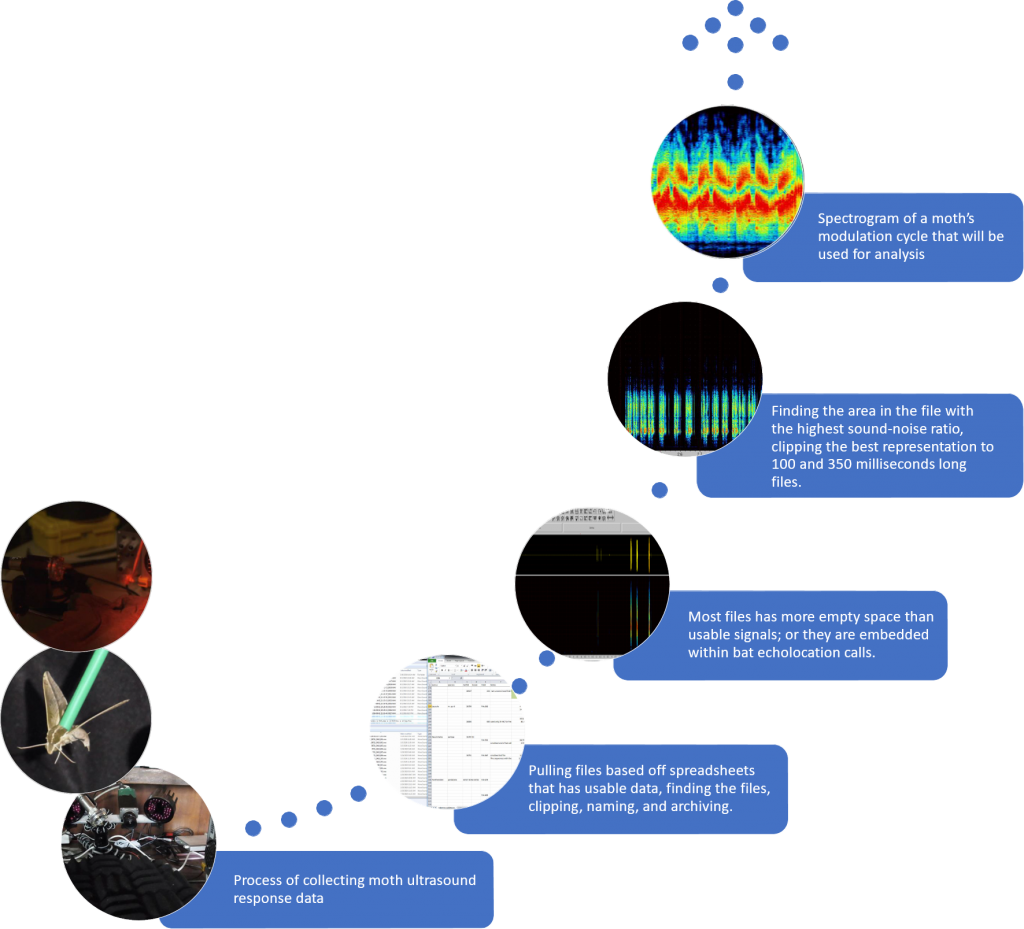
Results
Work in progress
Conclusions
We speculate that moths with similar clicking patterns will be clustered together. However, data archiving is still in progress.
Additional Information
For questions or comments about this research, contact Alyssa Culver at alyssaculver@u.boisestate.edu.